MC-3928
| Name | |||
|---|---|---|---|
| Unique ID | MC-3928 | ||
| Original ID | T-010 (Miyachi et al., 2021) | ||
| Common Name | |||
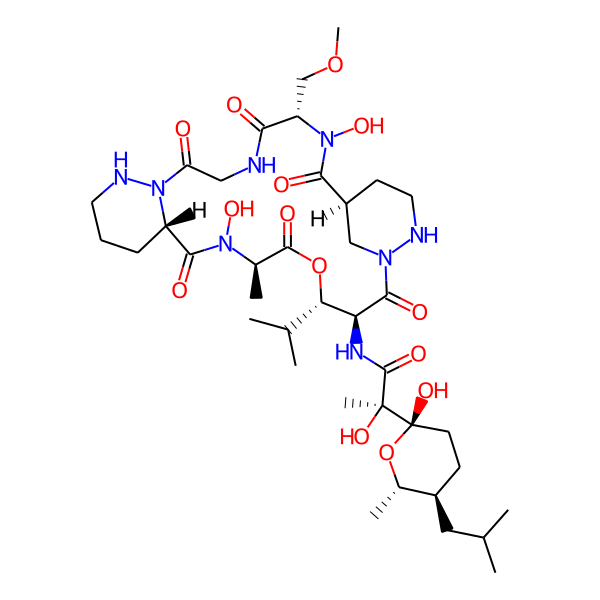
| Structure Representations | |||
| InchiKey | AMKPJJRZPMAWNG-MYHOCBOOSA-N | ||
| Isomeric SMILES | COC[C@H]1C(=O)NCC(=O)N2NCCC[C@H]2C(=O)N(O)[C@H](C)C(=O)O[C@@H](C(C)C)[C@H](NC(=O)[C@](C)(O)[C@]2(O)CC[C@@H](CC(C)C)[C@H](C)O2)C(=O)N2C[C@H](CCN2)C(=O)N1O | ||
| SMILES (Ring) | C1CNCCCOCCNCCNCCNCCNC1 | ||
| Permeability | |||
| Assay | PAMPA | ||
| Endpoint | Papp | ||
| Value | 7.4 | ||
| Unit | 10-6 cm/s | ||
| Standardized Value | -5.13 | ||
| Molecule Descriptors | |||
| MW (Da) | 856.97 | NRotB | 8 |
| HBA | 16 | Kier Index (Φ) | 16.40 |
| HBD | 8 | AR | 0.60 |
| cLogP | -1.84 | Fsp3 | 0.82 |
| TPSA (Å2) | 289.18 | MRS | 20 |
| Unique ID: ID for each unique molecule in this dataset; Original ID: The name or ID in original source; Standaridized value: Logarithmic values for permeability values (unit 10-6 cm/s), original data with sign '<' or '>' were removed during data standardization | |||
| Abbreviations: MW (Da): Molecular Weight; HBA: Hydrogen bond acceptor; HBD: Hydrogen bond donor; cLogP: Calculated lipophilicity; TPSA: Topological polar surface area; NRotB: Number of rotatable bonds; Φ: Kier flexibility Index: Fsp3: fraction of sp3 carbon atoms; MRS: macrocyclic ring size; AR: Amide ratio; | |||
Back to Browse